I am leading the research group Trustworthy Machine Learning at the Institute for Artificial Intelligence in Medicine (IKIM), located at the Ruhr-University Bochum.
My group offers Master’s thesis topics for students of the UA Ruhr (University Duisburg-Essen, Ruhr University Bochum, and TU Dortmund).
News
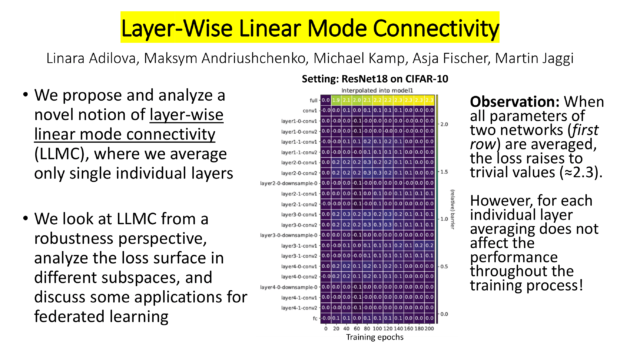
- My colleagues Linara Adilova, Maksym Andriushchenko, Asja Fischer, Martin Jaggi and I published a paper on Layer-wise Linear Mode Connectivity at ICLR 2024 (A*, top 7%).
- My colleagues Fan Yang, Pierre Le Bodic, Mario Boley and I published a paper on Orthogonal Gradient Boosting for Simpler Additive Rule Ensembles at AISTATS 2024 (A, top 16%).
- We presented 2 workshop papers at this year’s ICML in Honolulu: FAM: Relative Flatness Aware Minimization with Linara Adilova, Amr Abourayya, Jianning Li, Amin Dada, Henning Petzka, Jan Egger, and Jens Kleesiek at the Workshop on Topology, Algebra, and Geometry in Machine Learning and Informed Novelty Detection in Sequential Data by Per-Cluster Modeling with Linara Adilova and Siming Chen at the Workshop on Artificial Intelligence & Human-Computer Interaction.
- My colleagues Jianning Li, André Ferreira, Behrus Puladi, Victor Alves, Moon Kim, Felix Nensa, Jens Kleesiek, Seyed-Ahmad Ahmadi, Jan Egger, and I published a paper on “Open-source skull reconstruction with MONAI” in the Elsevier journal SoftwareX (IF: 2.87).
- My colleagues Jonas Fischer, Jilles Vreeken and I had our paper “Federated Learning for Small Datasets” accepted at ICLR 2023 (A*, top 7%).
- My colleagues Osman Mian, Jilles Vreeken and I had our paper “Information-Theoretic Causal Discovery and Intervention Detection over Multiple Environments” accepted at AAAI 2023 (A*, top 7%).
- My colleagues Osman Mian, David Kaltenpoth, Jilles Vreeken and I had our paper “Nothing but Regrets – Privacy-Preserving Federated Causal Discovery” accepted at AISTATS 2023 (A, top 16%).
- My colleagues Junhong Wang, Yun Li, Zhaoyu Zhou, Chengshun Wang, Yijie Hou, Li Zhang, Xiangyang Xue, Xiaolong Zhang, Siming Chen, and I published our paper “When, Where and How does it fail? A Spatial-temporal Visual Analytics Approach for Interpretable Object Detection in Autonomous Driving” in IEEE Transactions on Visualization and Computer Graphics (IF: 5.23).
Research Interests
I am interested in the theoretically sound application of machine learning to distributed data sources which entails four major challenges: The computational complexity of processing very large datasets, the often prohibitive communication required to centralize this data, the privacy-issues of sharing highly sensitive data, and the trustworthiness of the resulting model. My goal is to develop trustworthy machine learning methods. This means that ideally the learning method can be efficiently executed or parallelized and the resulting model is trustworthy in the sense that its performance can be guaranteed, it is robust against adversarial behavior, and its training preserves the privacy of sensitive data. I worked on in-situ methods that process data locally, thus using local processing power, reducing communication, and maintaining data privacy. I developed machine learning, online learning and online optimization algorithms for distributed and decentralized learning, both from batch data and data streams. As learning methods, I used (generalized) linear models, kernel methods, and deep learning. To ensure their trustworthiness, I analyzed them from a learning theory perspective to provide strong guarantees on their behavior and performance. I applied these approaches with partners in the industry on healthcare applications, but also on cybersecurity, material science, and autonomous driving.
Applying machine learning in healthcare poses novel and interesting challenges. Data privacy is paramount, applications require high confidence in model quality, and practitioners demand explainable and comprehensible models. In my research group on Trustworthy Machine Learning I tackle these challenges, investigating novel approaches to privacy-preserving federated learning, the theoretical foundations of deep learning, and the collaborative training of explainable models.
Curriculum Vitae – Highlights
Michael Kamp is the leader of the research group Trustworthy Machine Learning at the Institut für KI in der Medizin (IKIM), located at the Institute for Neuroinformatics at the Ruhr-University Bochum. In 2021 he was a postdoctoral researcher at the CISPA Helmholtz Center for Information Security in the Exploratory Data Analysis group of Jilles Vreeken. From 2019 to 2021 I was a postdoctoral research fellow in the Data Science & AI Department at Monash University and the Monash Data Futures Institute, where he is still an affiliated researcher. From 2011 to 2019 he was a data scientist at Fraunhofer IAIS, where he lead Fraunhofer’s part in the EU project DiSIEM, managing a small research team. Moreover, he was a project-specific consultant and researcher, e.g., for Volkswagen and DHL, and designed and gave industrial trainings. Since 2016 he was simultaneously a doctoral researcher at the University of Bonn, teaching graduate labs and seminars, and supervising Master’s and Bachelor’s theses. Before that, he worked for 10 years as a software developer. He is a member of the editorial board of the Springer journal Machine Learning and a member of the ELLIS society.